THE KEY WE CODED
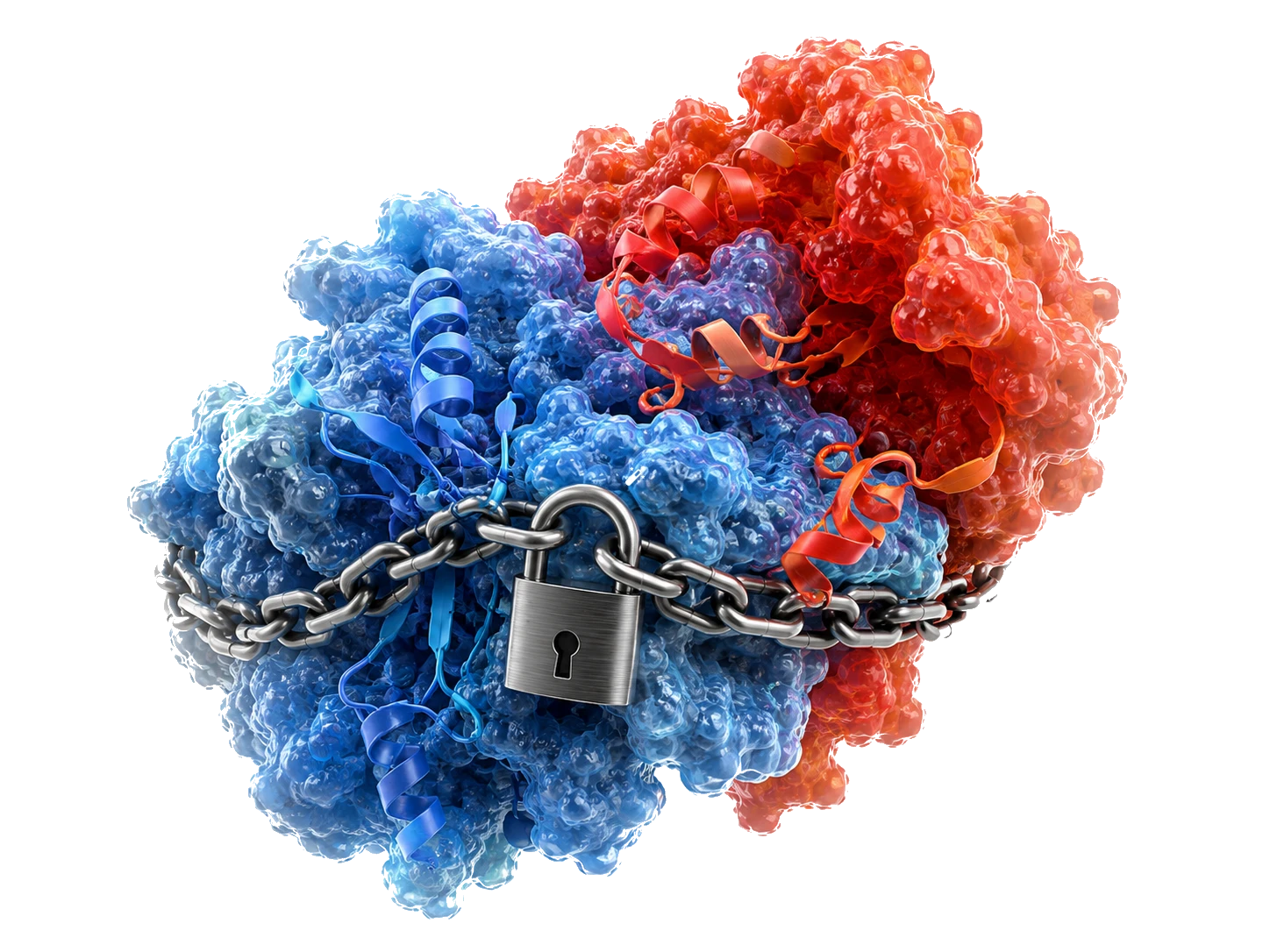
The Handcuffed Fighter
The human body already possesses a natural and highly effective tumor suppressor called the p53 protein. In half of all human cancers, the disease survives by overproducing an antagonist protein called MDM2, which physically binds to p53 and neutralizes it. This effectively holds the natural killer hostage, preventing the body from destroying the tumor. The body has its own cure, but cancer has it padlocked.

The Solution
Traditional drug discovery takes years and millions of dollars testing random chemicals, hoping one might be the key that fits the lock. We bypassed this slow, legacy model and used high-dimensional computational physics, math, and AI to design a custom molecular key called V37. In effect, we coded a key from scratch to target MDM2, pry it off p53, and reactivate the body's natural defense systems.
Breaking the Lab
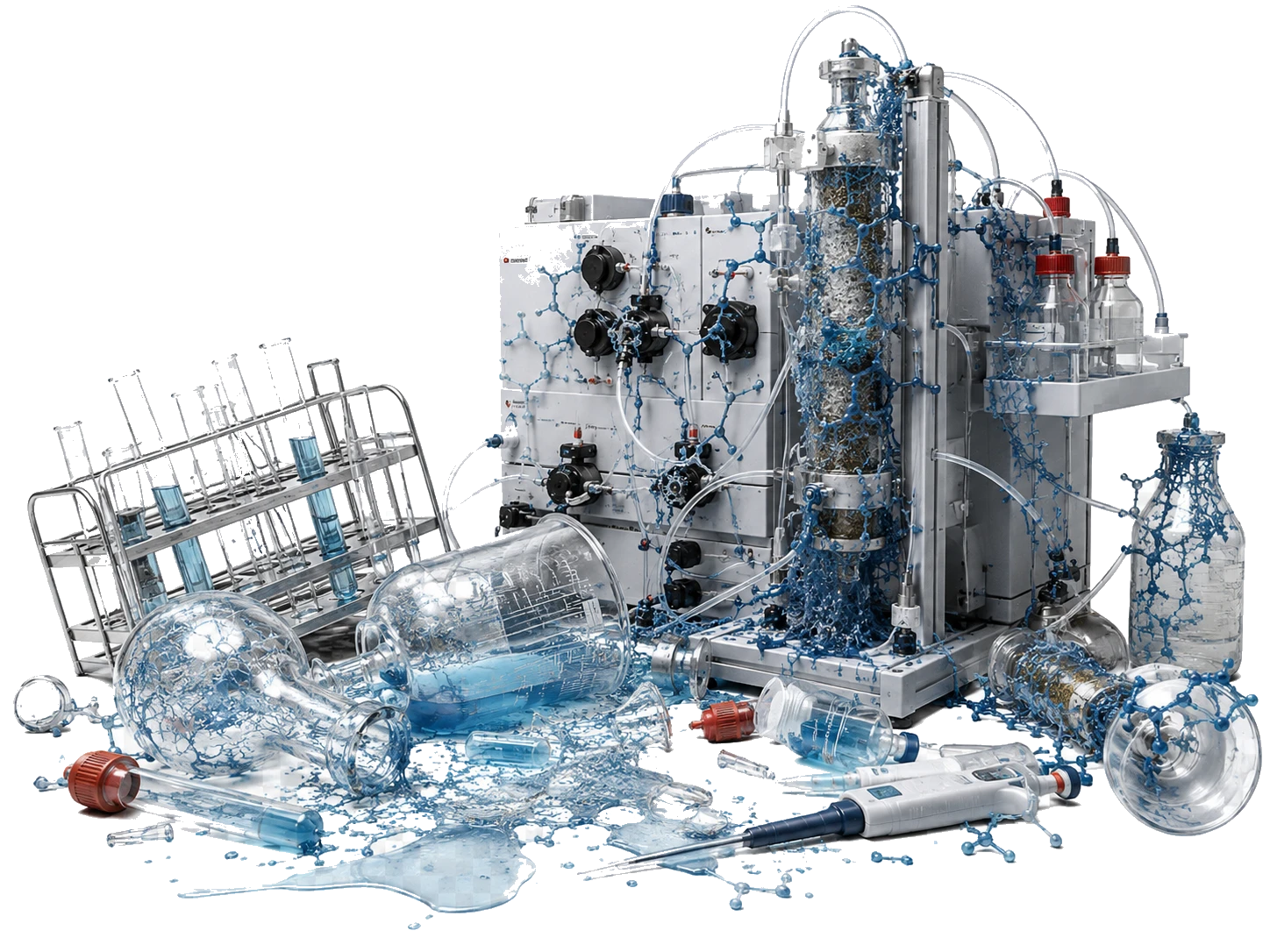
Because our molecular key is designed to bind to its target with extreme strength, it is highly dense and water-repelling. This caused the molecule to behave like an industrial adhesive inside standard automated manufacturing systems. It literally stuck to the inside of standard commercial purification machines. Standard automated factories simply could not process it.

The Solution
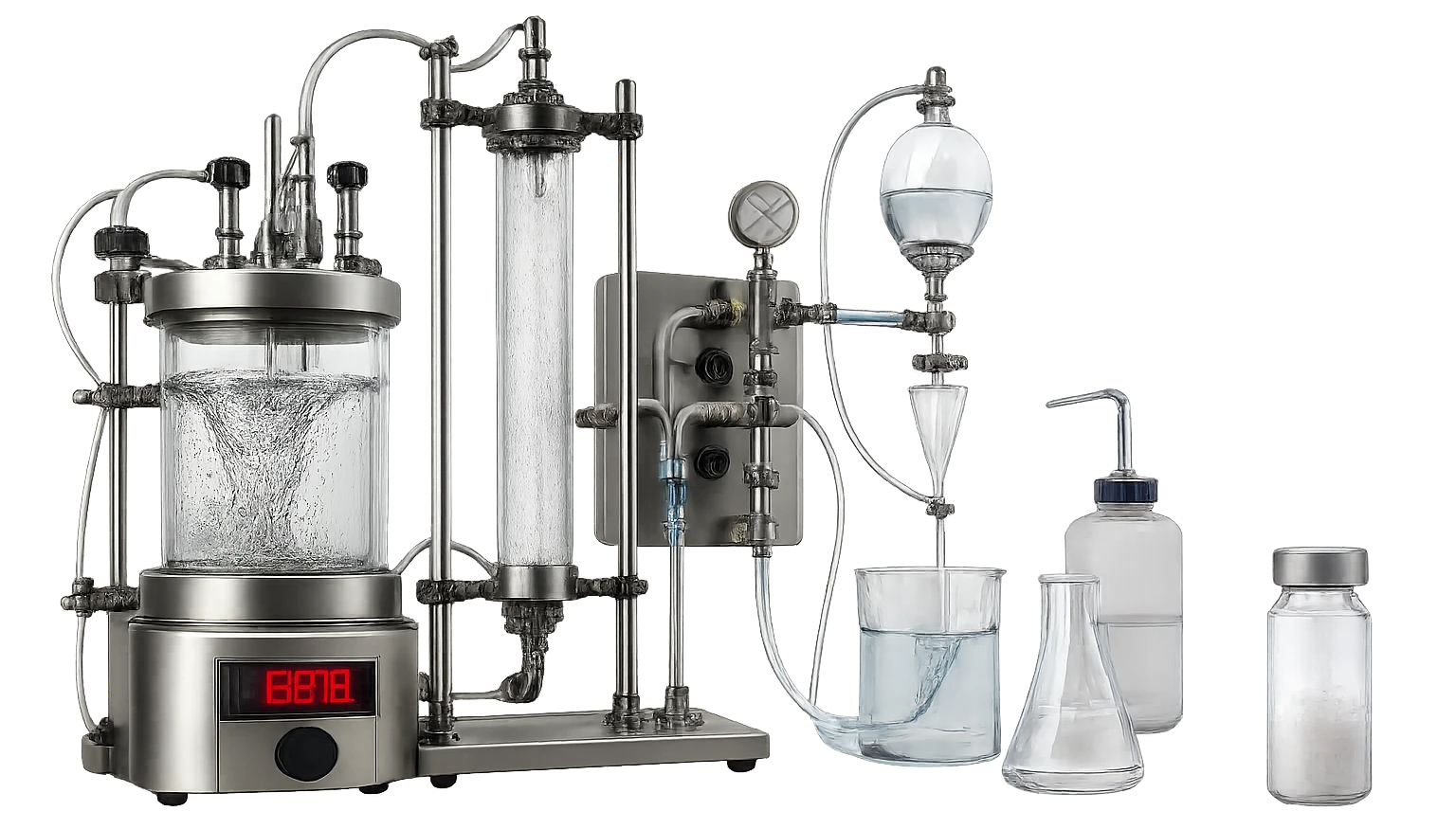
We solved the physical bottleneck. We designed a custom purification protocol using high heat (60 degrees Celsius) and specialized Isopropanol solvent washes to pry the molecule out of the machines. Through these custom engineering interventions, we successfully recovered four milligrams of the physical molecule. High-resolution mass spectrometers analyzed the resulting sample and confirmed the exact molecular weight of our key down to the decimal point: 1313.677. The math is now tangible matter.
